![]()
Organic chemists make judgements about how to represent molecular structures on a daily basis. When we give lectures, when we write papers, when we scribble on a napkin at lunch to share an idea with a colleague, we must make several choices (whether they be conscious or not). We must decide how much structural information to convey, from what angle to convey it, and whether to use simple lines or elaborate computer-generated graphics. Much has been written on these subjects (some by Roald Hoffmann), so for now I will just present a collection of images that I have produced for use in papers, in lectures, on journal covers, and just for fun.
Other examples of science related artwork can be found at the Art & Science Collaborations, Inc. website, as well as Nick DeMello's personal website.
A wonderful book of molecular images that predates the rise of computer graphics is The Architecture of Molecules, by Linus Pauling and Roger Hayward (W. H. Freeman & Co.: San Francisco, 1964).
Some of my non-chemistry images can be found here.
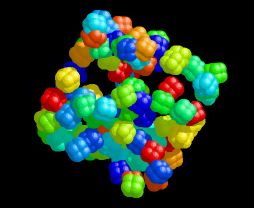
Cube of Cubanes
This image was created using a program called AutoDock that I used in my graduate research. It was designed for the automated docking of small ligand molecules into large macromolecular hosts. I discovered (by accident, of course) that if the box of space your ligand is allowed to sample is not centered on the macromolecule, you produce a random distribution of ligand positions inside a cube. I couldn't resist doing this again, this time on purpose and using my favorite cubic molecule. The resulting atomic coordinates were initially visualized with RasMol.
Of Mice
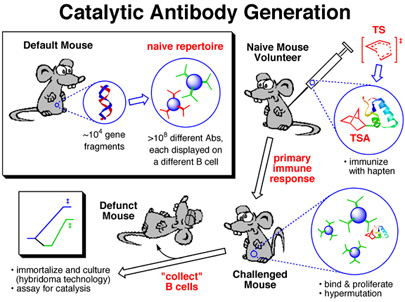
This image was created for use as a slide in several talks I've given on antibody catalysis. It was used to illustrate the key steps in the creation of catalytic antibodies. The image of duplex DNA in the top left panel was actually created from crystallographically determined coordinates using RasMol. The mice pictures are modified versions of an image I found somehwere on the web (unfortunately, I no longer remember where - my apologies to the creater of the original mouse image).
Boat (Full of Chemists) Conquers Chair
This image was created for the cover of the Journal of Organic Chemistry, but didn't quite make it there. It was to accompany a paper on intramolecular Diels-Alder (IMDA) reactions and was inspired by a call for creativity by my graduate advisor. The "chemists" (stylized versions of two of the paper's authors) are triumphantly hoisting their latest catch: a chairlike transition state for the IMDA reaction of a substituted hexadienyl acrylate.
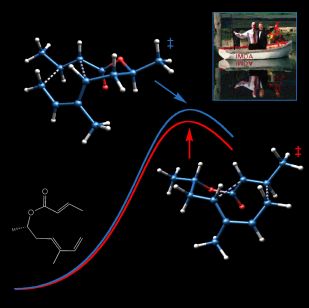
IMDATSs
This image is what resulted from the rejection of "Boat (Full of Chemists) Conquers Chair" by the editors of the Journal of Organic Chemistry (JOC). It depicts competing chairlike and boatlike transition states for the IMDA reaction of a substituted hexadienyl acrylate (shown as a line drawing) in the vicinity of a stylized pair of reaction coordinates. A miniature "Boat (Full of Chemists) Conquers Chair" is also present. The form in which this image actually appeared on the cover of JOC can be seen here. The transition state structures were visualized with SwissPdbViewer from computationally derived atomic coordinates.
Like Plastic
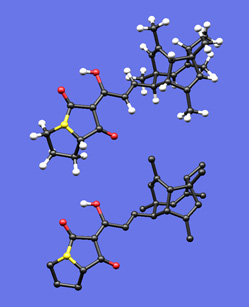
This image was created to accompany an essay I wrote on drawing polycyclic molecules. First I built a plastic model of the molecule and tried to photograph it, but when my photos came out blurry I thought I'd see if I could create an image with my Mac that actually looked like my plastic model. I built the molecule in a molecular modeling program, did a simple force-field minimization, took the resulting coordinates into SwissPdbViewer, and played with the settings (relative radii of balls and sticks, colors, bond lengths) until I felt the feel of my plastic model was captured in the rendered image. I was actually quite surprised how well this worked out -- so much so that someday I may write something on the benefits and dangers of this sort of representation. The bottom image is the same as the top, but with most of the hydrogen atoms removed.
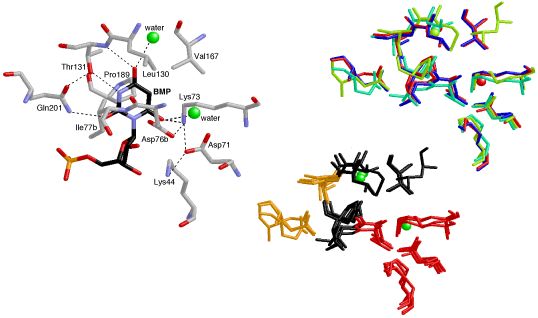
Conservation
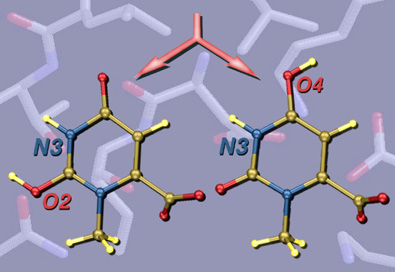
This image was created for a Highlight in ChemBioChem on the crystal structures and mechanisms of the world's most proficient enzyme: orotidine-5'-monophosphate decarboxylase (ODCase). The left panel shows a view of a typical ODCase active site with an inhibitor bound. The top right panel shows a superposition of the active sites from four independently derived x-ray crystal structures of ODCase•inhibitor complexes from four different species (each in a different color). The bottom right panel shows the same superposition now color-coded by residue-type: hydrophobic in black, polar but neutral in gold, charged in red, and waters in green. The molecular structures were visualized using RasMol.
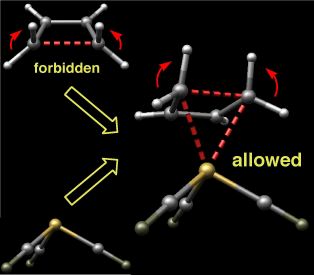
Demoniac Intervention
This image depicts the transition state for the disrotatory ring-opening of cyclobutene complexed to iron tricarbonyl. The disrotatory process in the absence of the organometallic fragment is symmetry forbidden, but upon complexation becomes the favored pathway. Woodward and Hoffmann once wrote:
"A very powerful Maxwell demon... could seize a molecule of cyclobutene at its methylene groups, and tear it open in a disrotatory fashion to obtain butadiene. But without such demoniac intervention, molecules will very rarely comport themselves in like fashion." (R. B. Woodward & Roald Hoffmann, Conservation of Orbital Symmetry, 1970)
In a sense, the iron tricarbonyl group here is providing just this sort of demoniac intervention. The molecular structures were visualized with SwissPdbViewer from computationally derived atomic coordinates.
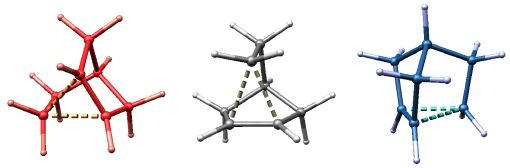
The Same or Not the Same?
The name of this image alludes to the title of a book written by my postdoctoral advisor (Roald Hoffmann, The Same and Not the Same, 1997). I leave it to you to figure out what these three structures represent. The molecular structures were visualized with SwissPdbViewer from computationally derived atomic coordinates.
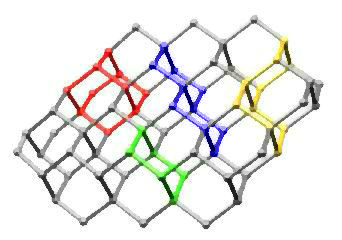
In Diamond
This image highlights chair cyclohexane, adamantane, trans-decalin, and cis-decalin substructures in a portion of the diamond lattice. It is based on an idea I had during graduate school for the cover of an undergraduate organic chemistry textbook. Maybe someday...
Tenuous
The molecule in this image (one designed by Roald Hoffmann years ago) contains two potentially aromatic systems (cyclopropenyl cation and cyclopentadienyl anion) connected by a methylene bridge. Computations may someday address whether these two care to conjoin through a covalent bond or prefer instead to remain in relative isolation. The molecular structure was previsualized with SwissPdbViewer from computationally derived atomic coordinates.
Unbuildable
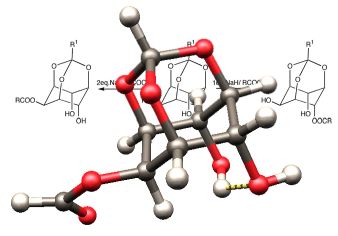
On occasion, despite the best efforts of authors, editors, and reviewers, a wonderful accident occurs and a drawing of an impossible (or at least highly improbable) molecular conformation slips into the primary research literature of organic chemistry. One such example is shown in this image. The inspiration for creating this image came from Ken Houk, who described these structures as "Escher molecules." The molecular structure in the foreground was previsualized with SwissPdbViewer from computationally derived atomic coordinates.
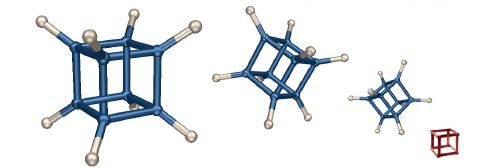
Cubain't
This image is my spin on the "impossible box" concept (an example is shown in red). The molecular structure of cubane was previsualized with SwissPdbViewer from computationally derived atomic coordinates.
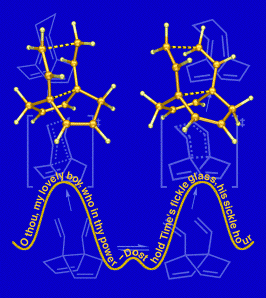
Fickled
This image was created to accompany a paper describing computations on molecules called "fickle hexadienes" and appeared on the cover of the March 8, 2002 issue of the Journal of Organic Chemistry. These molecules can rearrange in two stereochemically distinct ways (one chairlike and the other boatlike), but since the energies of the transition structures for both pathways are the same, the fickle hexadienes have no clear preference for one pathway over the other. The background of the image includes line drawings of the species involved in the rearrangement reactions, and the foreground consists of the two competing transition structures above a stylized potential energy surface adorned with a verse from Shakespeare. The transition state structures were visualized with SwissPdbViewer from computationally derived atomic coordinates. This image was also chosen to appear in the Journal of Organic Chemistry 2003 Calendar.
Models
This image of a ball and stick model of a tetrasubstituted methane is composed entirely of images from my collection of pictures of chemists posing with molecular models. It was inspired by a poster I saw in Mike Smith's office at UConn. Click on the image to zoom in.
Oh, Protonation
This image was created for Jeehiun Lee. Overlayed on a background of key residues from the active site of the enzyme ODCase are structures of differently protonated substrate models. The active site image was created from x-ray derived coordinates using RasMol, and the protonated substrate structures were created from computationally derived atomic coordinates using SwissPdbViewer.
Decarboxylation Machine
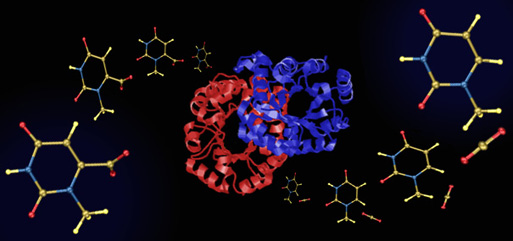
This image was also created for Jeehiun Lee. It consists of a central image of the enzyme ODCase flanked by cascading N-methylorotate molecules on the left and N-methyluracil and carbon dioxide molecules on the right - simple models of the natural substrate for ODCase (orotidine monophosphate) and the products of its decarboxylation. The enzyme image was created from x-ray derived atomic coordinates using RasMol, and the substrate and product structures were created from computationally derived atomic coordinates using SwissPdbViewer. This image appeared in the October 8, 2002 issue of Access. It is also included in the NSF Image Library.
U Exploded
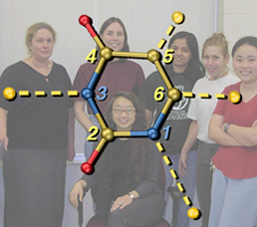
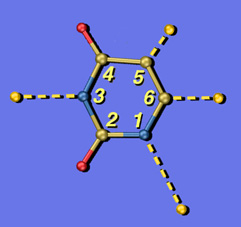
These are two versions of another image created for Jeehiun Lee. Each shows a uracil molecule with its four protons exploded off of the framework. Protons that are farther from the ring are more acidic than those closer to the ring. The background for the version on the left is a photo of the Lee group. The uracil structure was previsualized with SwissPdbViewer using computationally derived atomic coordinates.
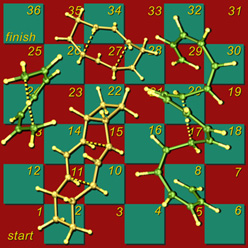
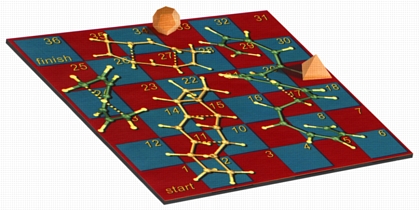
Snakes & Ladders
This image shows my version of the classic children's game Snakes and Ladders, using [1,7]-sigmatropic shiftamer transition structures as the snakes and [3,3]-sigmatropic shiftamer transition structures as the ladders. The structures were previsualized with SwissPdbViewer using computationally derived atomic coordinates. Below is a more recent variation with wooden polyhedra as game pieces. A version of this image appeared on the cover of an issue of Accounts of Chemical Research that featured an article on sigmatropic shiftamers.
The Circe Effect
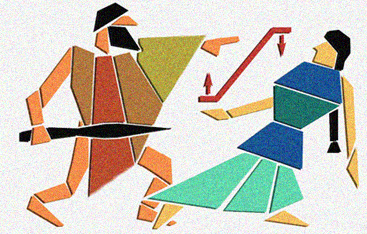

One day I was telling a friend about the enzyme ODCase. She thought I had made up the name as a play on "Odysseus." I thought that was rather funny, especially since one proposed mechanism for ODCase is based on something called the "Circe Effect." In short, this mechanism involves rate acceleration caused by selective destabilization of the reactant when complexed to the enzyme, rather than selective stabilization of the transition state. These two scenarios are shown schematically in red, with Circe favoring reactant destabilization and Odysseus favoring transition state stabilization. Perhaps you can guess which proposal I favor? The layout of this image was inspired by an ancient piece of art.
Full Circle
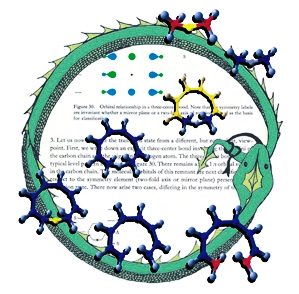
This image was created for the cover of an issue of Angewandte Chemie that featured an article on the geometric and electronic connections between different types of pericyclic reactions. The background is a page from Woodward and Hoffmann's classic Conservation of Orbital Symmetry. The molecular structures are transition structures for [1,8]-sigmatropic hydrogen shifts (center), 8-pi electrocyclization (bottom) and hydride transfer between carbenium ions (top); these structures were previsualized with SwissPdbViewer using computationally derived atomic coordinates. The circle is an orobouros - a mythical serpent eating its own tail - that I created in response to a suggestion by Roald Hoffmann; it is meant to emphasize both the cyclic nature of pericyclic transition structures as well as the fact that the article accompanying this illustration marked Roald's return to the topic of simple pericyclic reactions of hydrocarbons, nearly three decades after he first described many of their key electronic features.
Frame Shifting
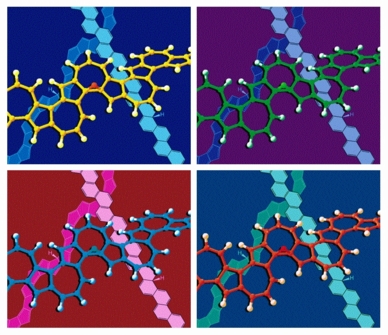
Each frame of this image shows a ball-and-stick model of a transition structure for the shift of a hydrogen atom over one face of a polyazulene polymer in front of line drawings of mono-hydrogenated polyphenacene and polypentalene polymers. A simplified version of this image appeared on the cover of an issue of the European Journal of Organic Chemistry that featured an article on sigmatropic shiftamers. The arrangement of the four frames was obviously inspired by some of the works of Andy Warhol. The structures were previsualized with SwissPdbViewer using computationally derived atomic coordinates.
Insensitivity
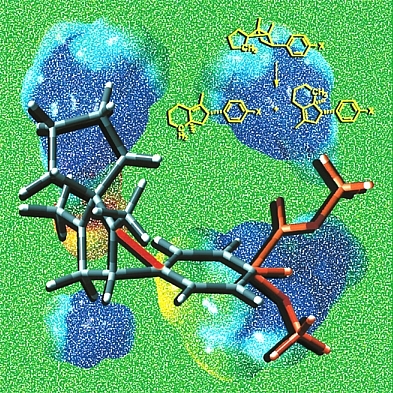
This image shows a model of a 1,3-diyl radical cation, an intermediate in the yellow reaction, stabilized by bridging of an appended aromatic ring. Several different substituents X are shown in orange at the bottom right of the image. Surprisingly, as demonstrated by our collaborator Dan Little, the regioselectivity of the reaction shown is actually insensitive to the nature of the substituent. The background images are based on computed electrostatic potential surfaces of 1,3-diyl radical cations involved in various ring-opening reactions of the type shown in yellow. Structures were previsualized with Gaussview using computationally derived atomic coordinates.
Disconnected
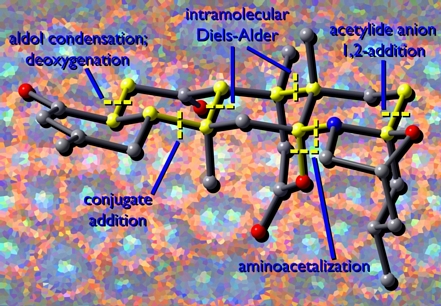
This image shows a model of a complex natural product called norzoanthamine. Synthetic disconnections are highlighted in yellow. The background image is derived from a photograph of the sea anemone (Zoanthus sp.) from which this molecule was first isolated. This image was created for Michael Nantz and George Zweifel based on an idea they had for the cover of their textbook, Modern Organic Synthesis, where a modified version of the image now appears. The structure was previsualized with Gaussview using computationally derived atomic coordinates.
Grubby
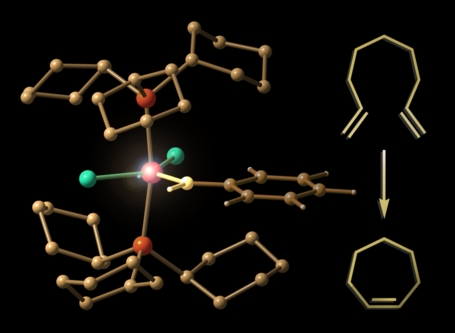
This image shows a model of Bob Grubbs' "first-generation" metathesis catalyst next to a representative example of an olefin metathesis reaction. This picture was created for use in advertising Prof. Grubbs' visit to UC Davis as part of the 2006 R. Bryan Miller Symposium. The structure was previsualized with Gaussview using computationally derived atomic coordinates.
Drugged Up
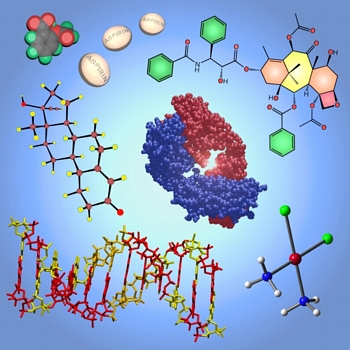
This image was created for use in advertising the Pharmaceutical Chemistry program launched at UC Davis in 2006. It shows various pharmaceuticals and pharmaceutical targets. These are (clockwise from top left): aspirin (space-filling model and pills), taxol (line drawing), the binding portion of an antibody (space-filling model), cisplatin (ball and stick model), a portion of DNA ("tube" model), and "the pill" (ball and stick model).
Royalty
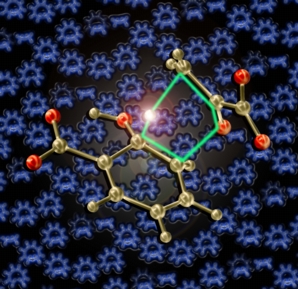
This image shows the transition state structure for a biological retro-ene reaction (computed by the Baldridge group) in front of transition structures for sigmatropic shifts and electrocyclizations morphing into each other and related carbocations. This picture was created for the cover of the Royal Society of Chemistry's Annual Reports on the Progress of Chemistry, Section B for 2006. The central transition structure was previsualized with Gaussview using computationally derived atomic coordinates.
Cavities
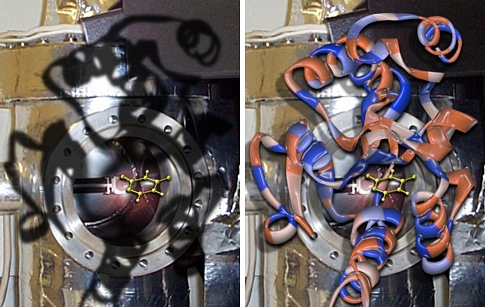

These images were created for Jeehiun Lee. Overlayed on a background of her mass spectrometer is the active site of the enzyme uracil DNA glycosylase. The yellow molecule in the center is deprotonated uracil, a species generated in the gas phase by Prof. Lee and also thought to be generated in the active site of the enzyme. The overlay of the spectrometer and enzyme "cavities" is meant to emphasize the ability of gas phase experiments to uncover fundamental properties of molecules that are relevant even to their reactivity in complex biological environs. The active site image was created from x-ray derived coordinates and the yellow molecule was previsualized with Gaussview using computationally derived atomic coordinates.
Mimics

This image shows a benzene ring interacting with the norbornyl cation via two different sorts of C-H---pi interactions (salmon and green). Like a mimic octopus (background), the norbornyl cation can change its appearance to suit its environment. Structures were previsualized with Gaussview using computationally derived atomic coordinates.
Analogous Pile
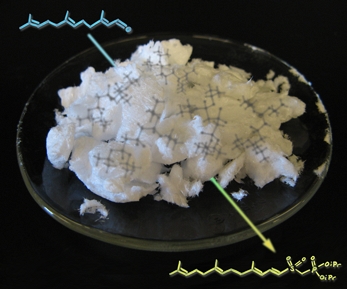

This image shows a reaction carried out in the lab by Mike. Crystals (made by Mike) of the reagent used to accomplish this transformation (developed by the Gervay-Hague group) are shown surrounding the arrow. Shadowy structures were previsualized with Gaussview using computationally derived atomic coordinates.
see BIG
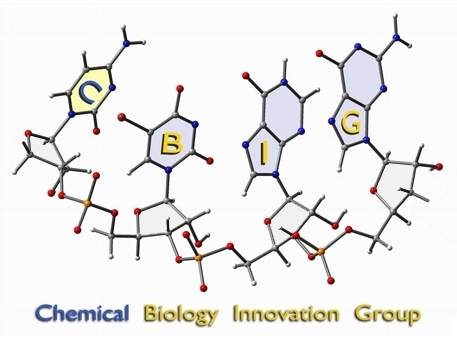
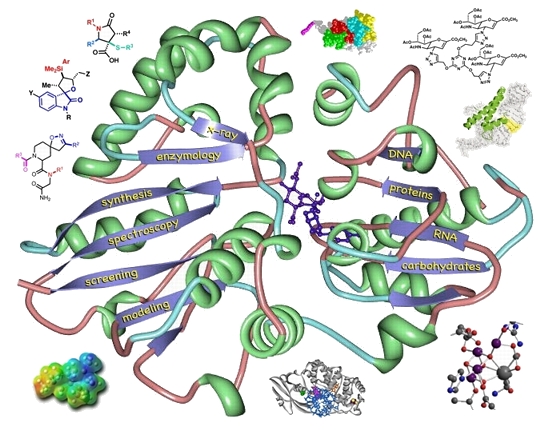
These two images were created for the CBIG program at UC Davis. The RNA sequence (cytidine - 5-bromo-uracil - inosine - guanine) was previsualized with Gaussview using a combination of x-ray and computationally derived atomic coordinates. The enzyme image was created from x-ray derived coordinates using the proteinWorkshop software available through the protein databank.
Brightness and Shadow

This image was created for Jeehiun Lee and Jeff Smiley for use on the cover of an issue of Organic and Biomolecular Chemistry that featured an article describing their research on the mechanism of the enzyme ODCase (shown as a shadow). In the center is shown the transition state structure for decarboxylation of orotidine-5'-monophosphate. Isotope effects for the two highlighted atoms were examined in the Lee-Smiley study. The enzyme image was created from x-ray derived coordinates and the transition state structure was previsualized with Gaussview using computationally derived atomic coordinates.
Into the Woods

This image was created to accompany an article on a diterpene-forming carbocation rearrangement that occurs on a bifurcating potential energy surface. In the center is shown a transition state structure for proton transfer (green) that occurs along a pathway that subsequently bifurcates towards to two structures shown in the distance. The structures were previsualized with Gaussview using computationally derived atomic coordinates.
Olives and Oil
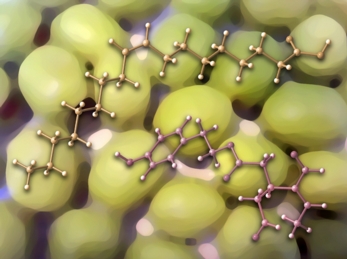

This image was created for colleagues in UC Davis' Department of Food Science and Technology. The two molecules shown, oleic acid and oleocanthal, are both present in olive oil. The structures were previsualized with Gaussview using computationally derived atomic coordinates. The olives in the background were found in Dean's fridge.
Kendall the Kid

This image was created (in partnership with Dean's UC Davis colleague Gang-yu Liu) to honor Dean's PhD advisor - Ken Houk - at the Mesilla Chemistry Workshop on Aromatic Interactions in Chemistry and Biology. Note that Mesilla is the location where Billy the Kid was put on trial.
Natural Barriers

This image was created for use on the cover of an issue of Natural Product Reports that featured a review article describing calculations on terpene-forming carbocation rearrangements. The background image is a photo taken by Dean at Yosemite in March, 2011. Attached to the fallen tree as branches are several computed transition state structures. A computed reaction coordinate floats on the surface of the foreground water. The transition state structures were previsualized with Gaussview using computationally derived atomic coordinates. The reaction coordinate is derived from an actual IRC calculation.
Think Pizza

This image was created to accompany a C&EN blog post by Selina Wang.
Spaceballs
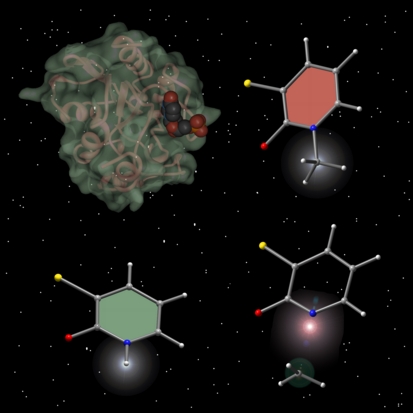
This image was created for Jeehiun Lee for use on the cover of an issue of the Journal of Organic Chemistry that featured an article describing her research on the differences between basicity and leaving group ability. At the top left is shown the crystal structure of uracil DNA glycosylase, the inspiration for this work. The enzyme image was created from x-ray derived coordinates using Protein Viewer via the Protein Databank and the other molecules were previsualized with Gaussview using computationally derived atomic coordinates.
Working with Your Hands
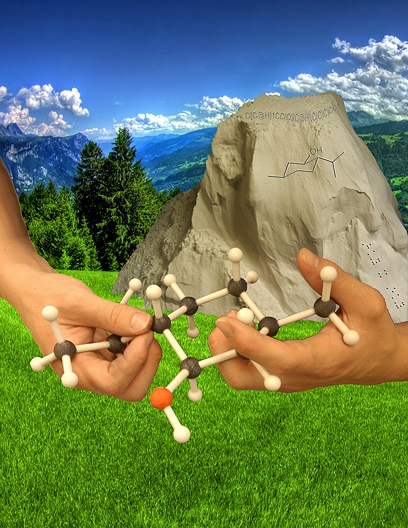

This image was created to accompany an article on our efforts to make computational chemistry research accessible to blind and visually impaired scientists. The model was printed on our 3D printer. The mountain is a pile of powder from which such models must be excavated at the conclusion of the printing process. Embossed on the mountain is a line drawing of the modeled structure, a SMILES string indicating its connectivity and a message in Braille.
From the Shadows
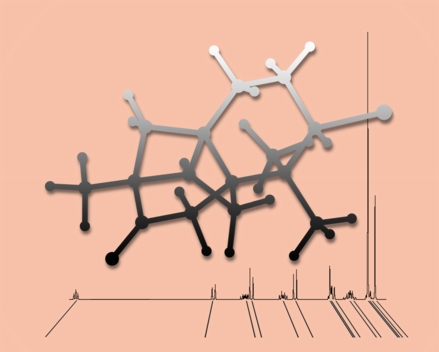
This image was created to accompany our review article on NMR chemical shift calculations. The shadowy molecule is aplydactone, floating in front of an experimental NMR spectrum and lines indicating values of computed chemical shifts.
Siege

This image was created for the cover of an issue of Organic and Biomolecular Chemistry that featured an article describing our research on ladderanes. This article is about how breaking open ladderane lipids may prevent small toxic molecules from passing through cellular membranes. This concept is represented here using ladderanes as medieval siege ladders. Note that the lefthand pennant is is emblazoned with UC Davis colors and the righthand pennant is a stylized version of the Tantillo group logo. Molecules were previsualized with Gaussview using computationally derived atomic coordinates.
Shiny Cubes
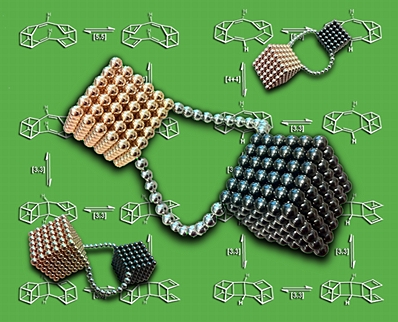

This image was created for the inside cover of an issue of Phys Chem Chem Phys that featured an article describing our research on sigmatropic shifts of coupled cubanes.
Planting a Seed
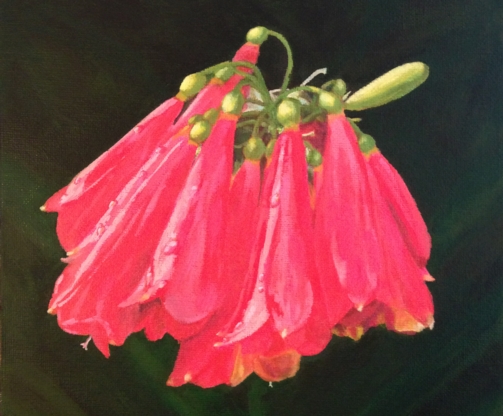

This image was created for the cover of an issue of Natural Product Reports that featured an article on applications of quantum chemistry to problems in natural products chemistry. The image is a photograph of a painting by Dean of Clivia nobilis, a plant from which a natural product (nobilistitine), whose structure we reassigned using NMR computations, was isolated.
Accessibility
This image was created as a logo for Accessible Science - do you see the message in the molecule (which was rendered so as to be readable using one's hands when printed large enough)?
Swarthy

This image was created to accompany an article on applications of quantum chemistry to problems in RNA chemistry. The model was printed with our 3D printer and the background shows our in-house computer cluster.
In the Background
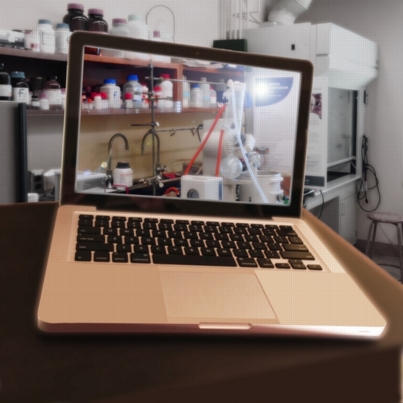

This image was created to accompany an article on applications of quantum chemistry to problems in synthetic chemistry.
Bubbles

This image was created to accompany an article on molecules designed to fight cystic fibrosis.
Flying Away
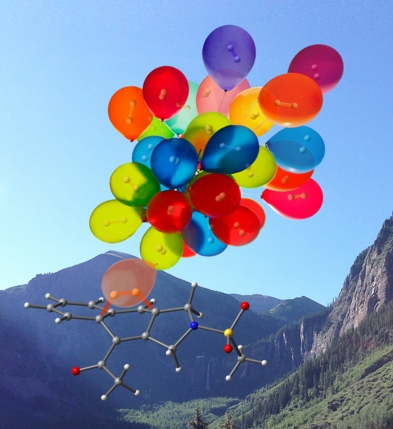

This image was created to accompany an article on the mechanism of a hydrogen-producing dehydro-Diels-Alder reaction (transition state structure for H2 explusion shown). The background is Telluride, CO. This image eventually ended up on the cover of Dean's book.
Detour
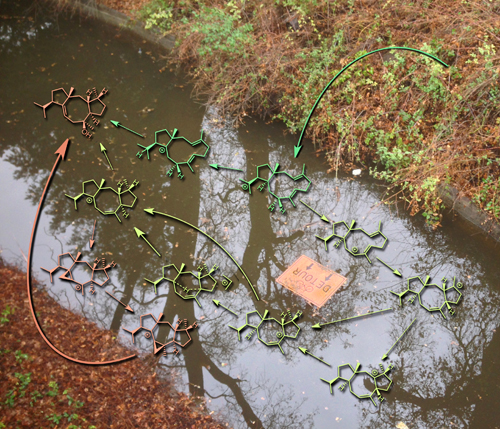

This image was created to accompany an article on the mechanism of a terpene-forming carbocation cyclization/rearrangement. The background photo was taken on the UCD campus.
Map
This image was created to accompany a special issue of Accounts of Chemical Research. The background consists of stylized images of computed transition state structures (created from images in ACS-published papers written by authors of articles for the special issue). The foreground consists of a “network diagram” where the relative positions of the nodes correspond roughly to locations of article authors on a world map (at a non-standard angle).
Haystack

This image was created to accompany an article on a newdocking approach for terpene synthases. The image used by the publisher highlighted one of the hidden carbocations.
Shall We Play a Game?

This image was created to accompany an article on a polycyclization reaction leading to dodecahedrane. The image was created from scratch and, yes, that little one is who you think it is!
Hanging Around...
This image was created to accompany an article on synthetic methods to make molecules related to some found in wine. The image was created from scratch and the background image started as a photo taken in Davis.
Clowns to the Left of Me...
This image was created to accompany an article on predicting chemical shifts for cyclic peptides. The image was created from scratch and the background image started as a photo taken in Chile.
Clowns to the Left of Me...

This image was created to accompany an article on a terpene-forming carbocation cyclization/rearrangement reaction that involves a post-transition state bifurcation. The image was created from scratch and the background image started as a photo taken in Ottawa.
One Way or Another...
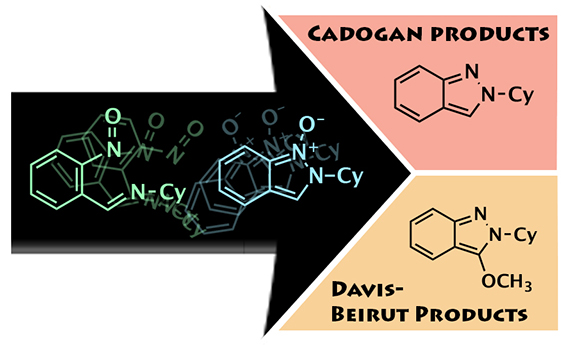
This image was created to accompany an article on a collaborative research project with the Kurth group. The image was created from scratch .
Hammer Time...
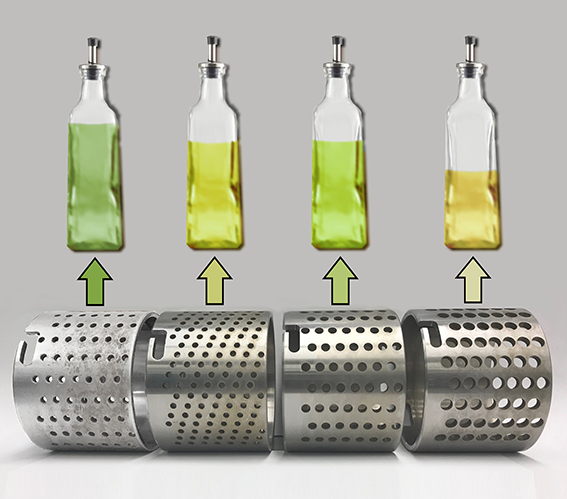

This image was created to accompany an article by alumna Selina Wang on crushing olives to make olive oil. The image was created from scratch and the background image started as a photo taken by Juan Polari of Selina's group.
Something Fishy...
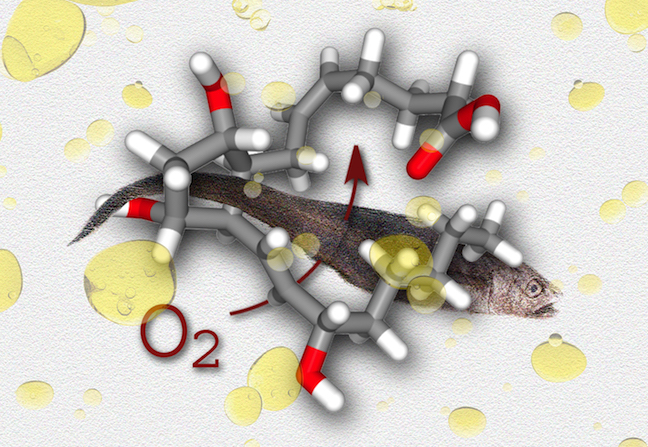
This image was created to accompany an article by alumna Selina Wang on fish oil.
It's What's on the Inside...
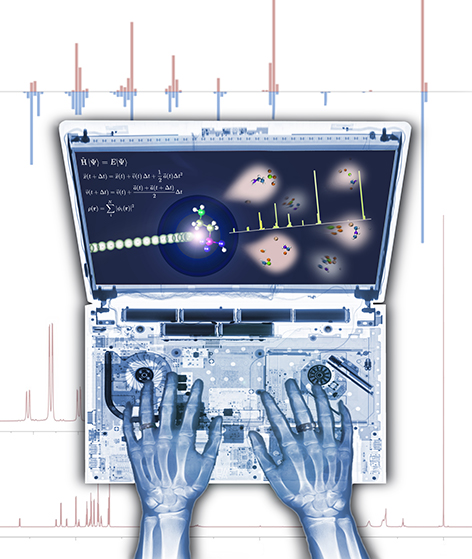
This image was created to accompany a review article on quantum chemical tools for metabolomics. We hoped it would be used as cover art, but, alas, it was not. A version of it was used, though, as a TOC graphic.
Chopped...
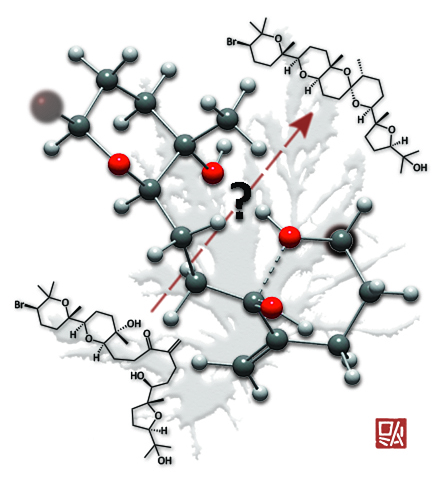
This image was created to accompany an article on the biosynthesis of polycyclic ether natural products. It was used both as the TOC graphic and, without my chop, as a supplementary cover. The background is the alga from which the specific natural products studied were isolated.
Decision Points...

This image was created to accompany an essay on discomfort in science. As such, it appeared on the cover of J. Phys. Org. Chem.
Dean's Driveway...
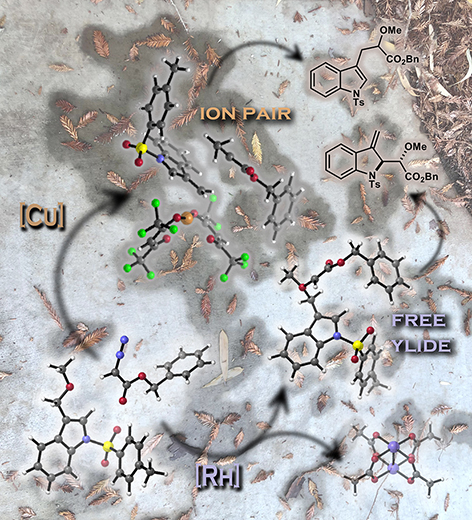
This image was created to accompany a paper involving ion pairs that react before equilibrating.
Dean's Kitchen...

This image was created to accompany a special issue of JOC on solvation effects in organic chemistry. It all started with a photo of a bowl of water taken at Dean's home...
Attached...
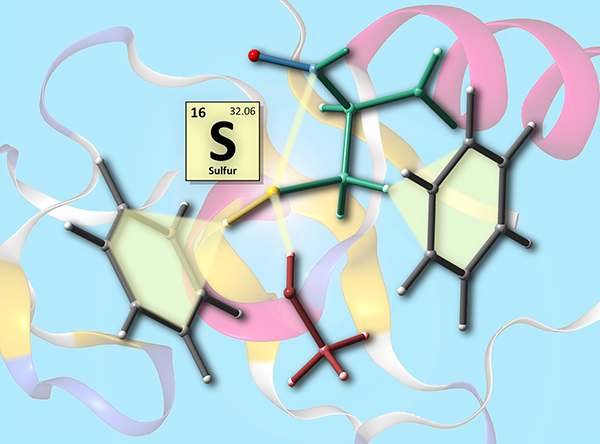
This image was created to accompany a review on the relevance of noncovalent interactions involving sulfur atoms to protein structure. It appeared on the cover of Org. Biomol.Chem.
Gilopus...

This image was created to accompany a paper on an organometallic reaction. It was generated by an AI told to make something like an octopus and seeded with an image from the paper showing an overlay of multiple catalyst conformations. The resulting image was then tuned up and the bakground and molecular structures added. It appeared on the cover of Chem. Eur. J.
Woodblocking...
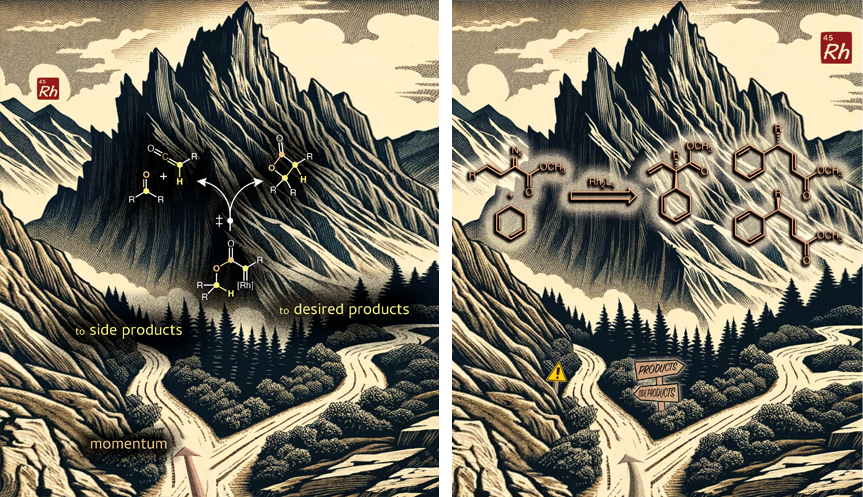
These images were created to accompany two papers on an organometallic reactions with PTSBs. The backgrounds were generated by an AI. The resulting images were then modified and molecular structures added. One image appeared on the cover of Acc. Chem. Res.
Bundt...

This image was created to accompany a paper on a fascinating old carbocation. Someone you might know actually baked that cake ;-) This image appeared on the cover of Chem. Sci.
Zwitty...
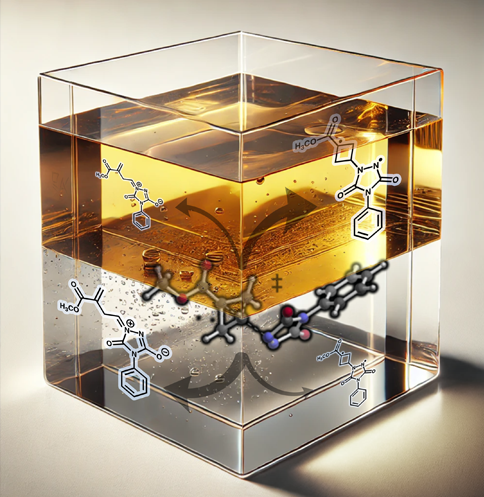
This image was created to accompany a paper on a PTSB with a product ratio that is sensitive to solvent polarity. The background was generated by an AI and modified. It didn't make it to the cover, but a version appeared as a TOC graphic.
Snake's Got Your Tail...
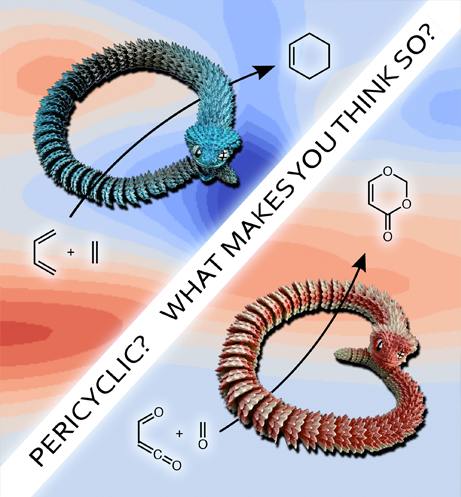
This image was created for a JOC cover accompanying a paper on a computational tool, PeriRxn, that may or may not be useful for determining whether a reaction is truly pericyclic. The background is derived from output of PeriRxn for a reaction that is (top left) and is not (bottom right) pericyclic. The key is that the pericyclic reaction involves continuous cyclic electron delocalization (snake grabs its tail) while the non-pericyclic reaction involves a break in delocalization (snake's head is near tale but not grabbing it).
Conjunction Injunction...
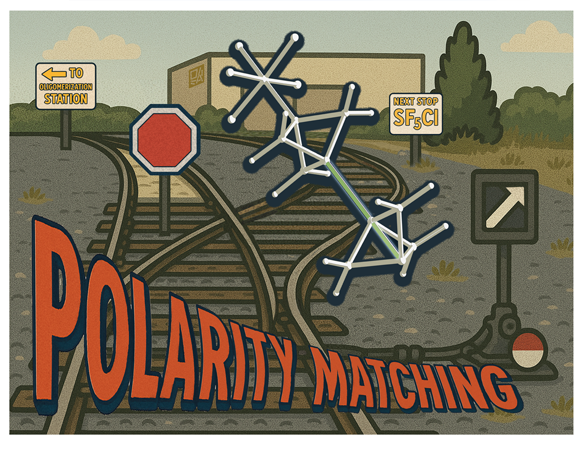
This image was created for a journal cover accompanying a paper with Cody Pitts' group, but it was rejected by the journal because I initially used AI to generate a draft of the background image, which was then heavily modified– unbeknownst to us, they had a zero AI policy. I still like the old-timey postcard feel though.